|
Likelihood
|
Conjugate prior
|
Posterior
|
|
Gaussian
|
Gaussian
|
Gaussian
|
|
Binomial
|
Beta
|
Beta
|
|
Poisson
|
Gamma
|
Gamma
|
|
Multinomial
|
Dirichlet
|
Dirichlet
|
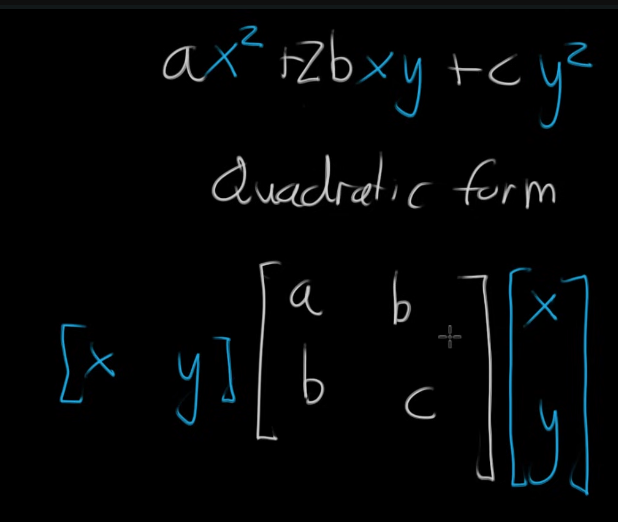
|
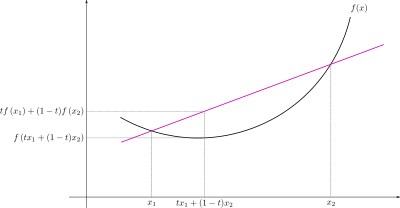
Function
|
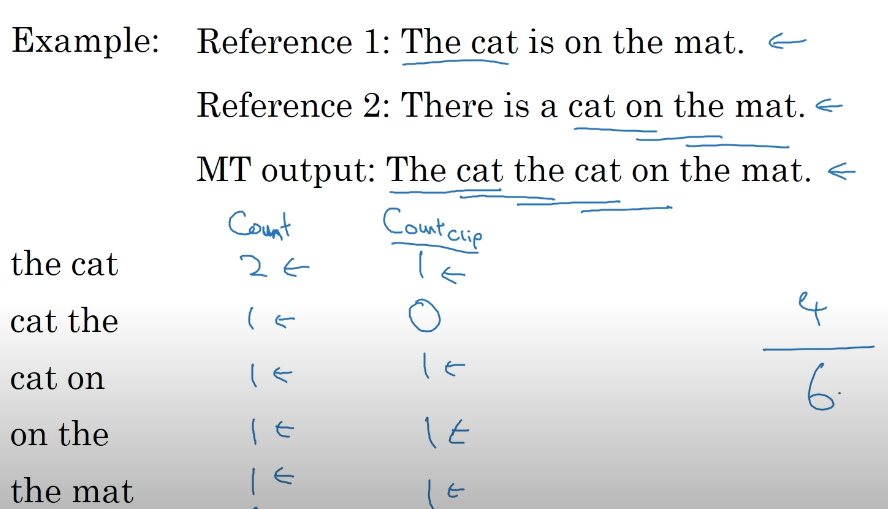
Example
|
1st derivative
|
2nd derivative
|
|
Loss function
|
Gradient
|
Hessian (symmetric )
|
|
|
NN linear layer
|
Jacobian
|
(higher order tensor)
|
|
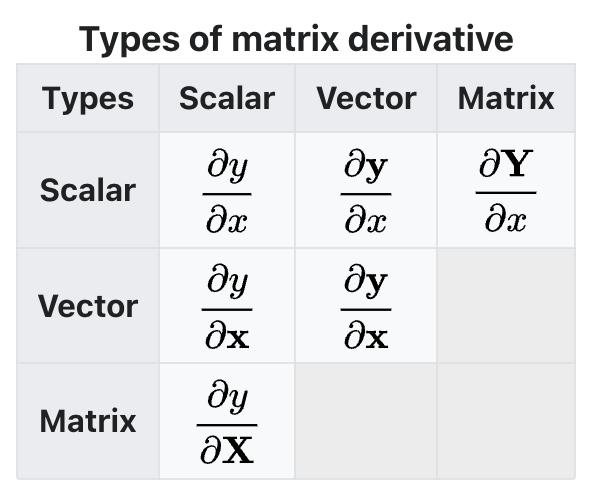
Name
|
Conditions
|
Decomposition
|
||
|
LU
|
Square matrix
|
, is lower tri, is upper tri.
|
, based on # row swaps
|
|
|
QR
|
Any matrix
|
, is ortho, is upper tri. Or , is diag.
|
||
|
Cholesky
|
PD symmetric matrix
|
where is lower tri.
|
||
|
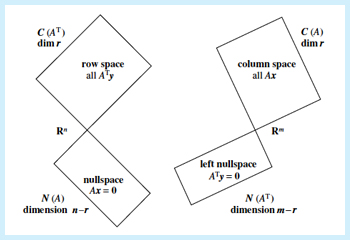
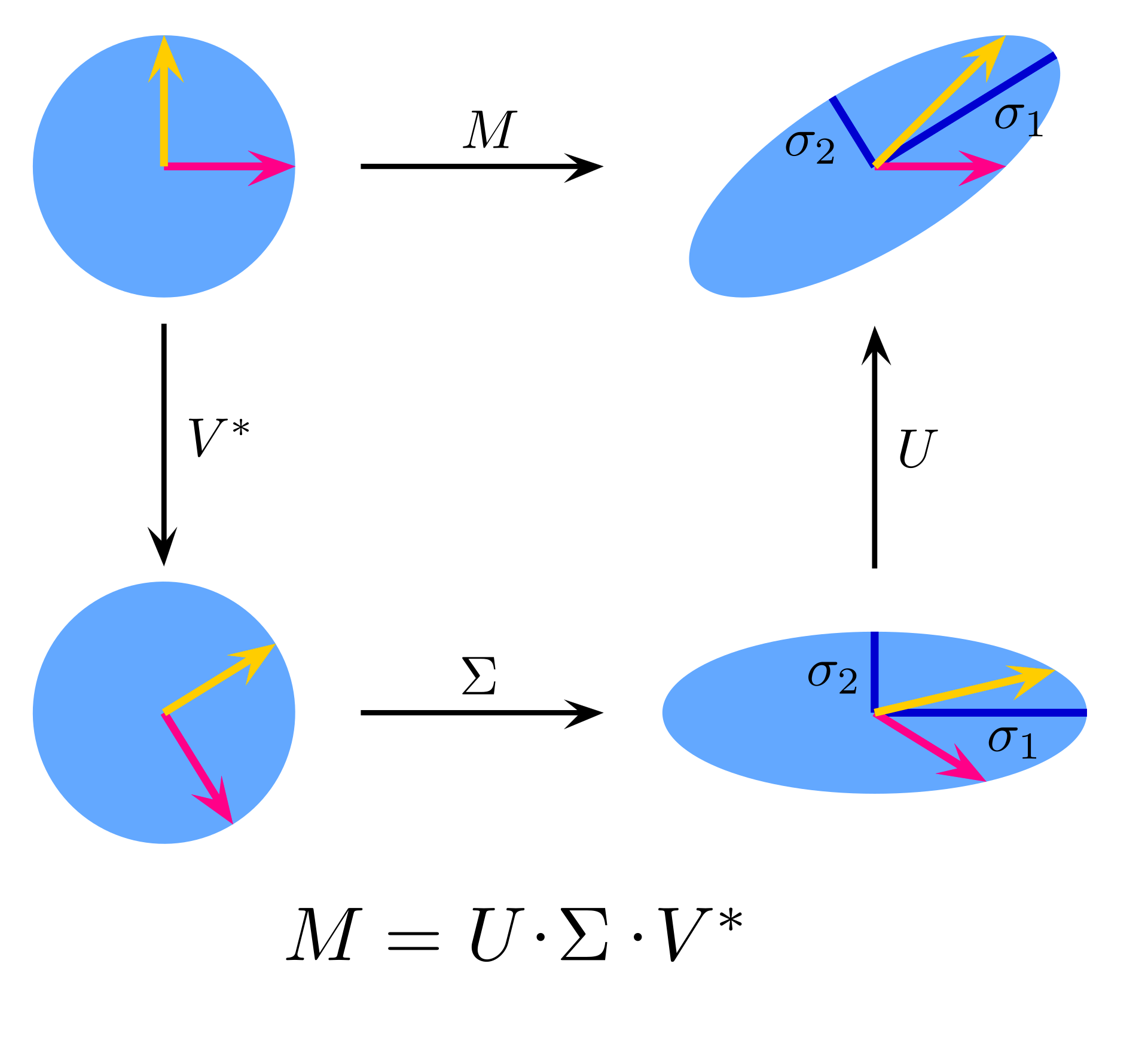
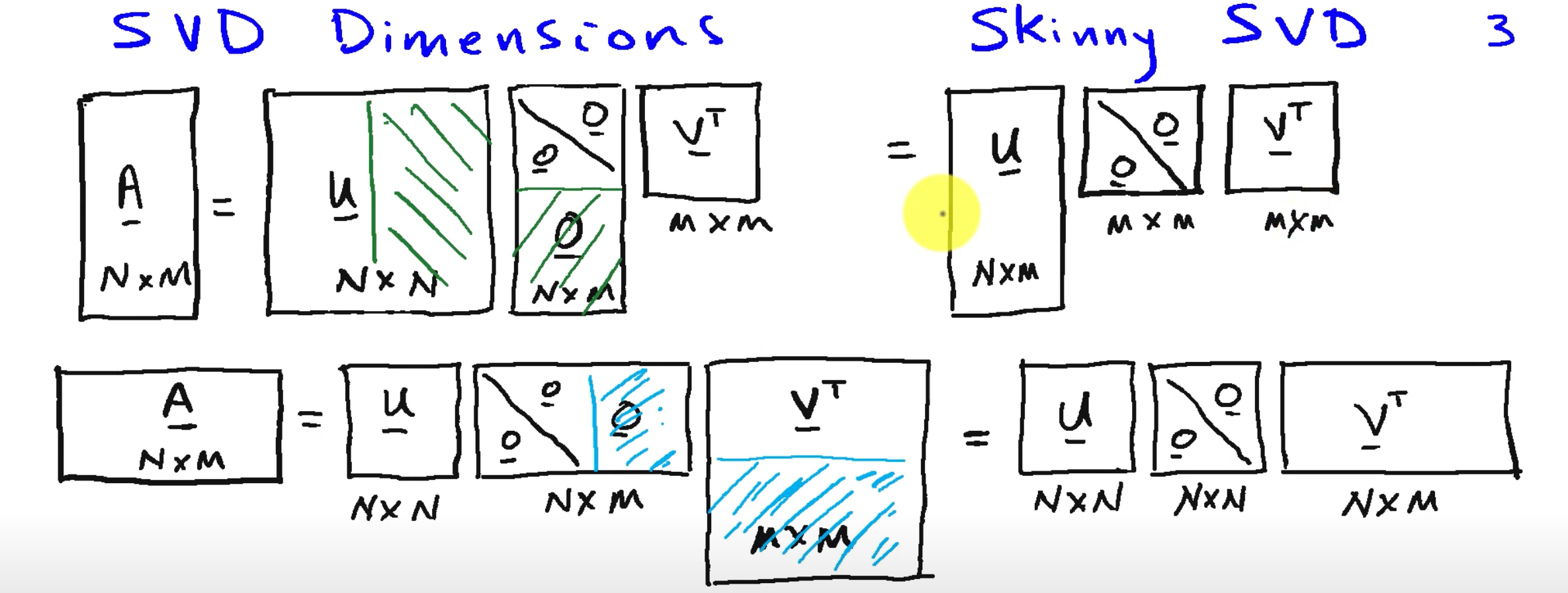
SVD
|
Any matrix
|
, is diag (ordered singular values), ortho.
|
||
|
EVD / Spectral
|
Square matrix
|
, is usually ortho (eigenvectors), is diag (eigenvalues).
|
||
|
NMF
|
Non-neg matrix
|
, also non-neg (and usu. lower rank approx)
|
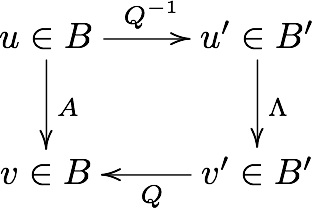

# Create a matrix A = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]]) # Compute the singular value decomposition (SVD) of the matrix U, S, Vh = np.linalg.svd(A) # Truncate the SVD to a lower rank k = 2 U_k = U[:, :k] S_k = S[:k] Vh_k = Vh[:k] # Reconstruct the matrix from the truncated SVD A_approx = U_k @ np.diag(S_k) @ Vh_k